To detect the tyrosine phosphorylated EpoR, phosphotyrosyl proteins from 500g of total cell protein have been immunoprecipitated with mAb 4G10 in IP buffer Tween20, 10g ml aprotinin, 1 mM PMSF, 5g ml antipain, 0. 5g ml leupeptin, 0. 7g ml pepstatin A, 1 Comprehensive inhibi tors, 3 mM DTT, 1 mM NaF, 1 mM Na3VO4 overnight in the cold. Soon after SDS Web page and membrane transfer, EpoR was detected with polyclonal anti EpoR and ECL. PI3 kinase assay 500g of total cell protein extract had been immunoprecipi tated in NP 40 lysis buffer with 4g anti phosphotyrosine mAb or with 4g PI3K p110 polyclonal anti physique, Santa Cruz respectively. Immu noprecipitates had been washed 3 instances with NP 40 lysis buffer, twice with 50 mM TrisHCl pH 7. 5 plus 500 mM lithium chloride, as soon as with 20 mM TrisHCl pH 7.
five with one hundred mM NaCl and 1 mM EDTA and after with 20 mM HEPES pH 7. five. 10g of lyophilised phosphatidyli nositol had been selleck then mixed with ten l of lipid kinase buffer and sonicated for 6 5 sec at 30% energy output with an MS72 sonotrode, chilling PI on ice amongst sonications. Sonicated PI was then mixed with 30 l LKB and added towards the immunoprecipitates. After 5 min on ice, 10 l LKB with 10 Ci 32P ATP and 2 mM magnesium chlo ride were added and samples incubated at 37 C for 15 min. The reaction was stopped with 200 l of 1 N hydrochloric acid. Just after extraction of the samples with 200 l of methanol chloroform, 50 l on the organic phase have been made use of for thin layer chromatography on LK6D plates. Running buffer was methanol chlo roform H2O NH4OH. TLC plates were analyzed on a Storm860 phosphoimager with ImageQuant 5. 2.
To confirm equal PI3K precipitation independent of cell stimulation, parallel samples of immunoprecipitates from protein extracts ana lyzed in lipid kinase assay had been separated by SDS Web page and blots probed with anti p110 mAb followed by ECL. Raf MEK coupled kinase assay The coupled kinase assay was accomplished basically as described with slight modifications. 500g of total cell protein extract were epigallocatechin immunoprecipitated with 4g anti c Raf1 or with 4g anti B Raf in buffer A TritonX 100, 10% glycerol. Immunoprecipitates had been washed three times with buffer A and twice with buffer B. 0. 2g acti vatable GST MEK1 with each other with five l buffer C and 10 l buffer B were added to the washed immunoprecipitates, samples have been mixed and incubated at 30 C for 20 min. The reaction was then disrupted by transferring samples into ice and adding 100 l buffer D. 25 l of this reaction mix was subsequently incubated with 20g kinase inactive GST Erk1K63M, two l buffer E and five Ci 32P ATP at 30 C for 15 min. The reaction was stopped by adding SDS Page sample buffer. Just after SDS Page, Raf activity was determined by detecting GST Erk1K63M phosphorylation with Image Quant five.
Monthly Archives: August 2014
To figure out specific involvement on the ERK1 2 activation in SS
To figure out distinct involvement of your ERK1 2 activation in SS RBC mem brane protein phosphorylation, each and every population of SS and AA RBCs was either treated or not treated having a potent MEK1 2 inhibitor, U0126, which specif ically inhibits ERK1 2 kinase activity. RBC membrane ghosts prepared from the resulting four populations of RBCs, were then either subsequently co incubated in the presence or absence of exogenous recombinant ac tive ERK2. Proteolytically digested mem brane fractions from every single of these eight exclusive samples have been then subjected to a previously described label absolutely free quantitative phosphoproteomics workflow utilizing re producible TiO2 phosphopeptide enrichments followed by selected ion chromatographic peak quantitation of accurate mass retention time aligned LC MS MS data to enable direct quantitative comparisons to become created across all therapy groups.
To minimize total analysis time, each and every sample was analyzed in analytical triplicate by a one dimensional LC MS MS analysis without having any added fractionation prior to TiO2 enrichment. Across all samples, 375 distinctive phosphopeptides corresponding to 155 phosphoproteins were identified at a peptide spectral hop over to this website false discovery price of 1. 0%. As localization of particular phosphorylated residues is essential for defining kinase distinct events, all phosphopeptides were subjected to ModLoc, a probability primarily based localization tool implemen ted within Rosetta Elucidator determined by the AScore algorithm. Approxi mately 74% of phosphorylated residues had Mod Loc scores above 15, and 66% had ModLoc scores above 20.
To assess the quantitative robustness of the label cost-free approach, the average technical coefficient of variation of retention time aligned phosphory lated peptide intensities of triplicate measurements within a therapy group had been calculated. i was reading this The imply %CV across all 375 phosphopeptides was 19. 8%, with 80% from the signals obtaining a %CVs much less than 27. 1%. The intensity on the phosphorylated peptide V173 R191 inside the ac tive site of ERK1 2 was applied to assess inter therapy group variation, which includes variation from TiO2 phospho peptide enrichment, as activated ERK2 was spiked in equal amounts to four of the eight samples. The typical %CV of this phosphopeptide inside any treatment group was 7. 0%, and across all ERK2 spiked samples was 18. 1%.
Constant using a majority of TiO2 enrichment primarily based worldwide mammalian phosphoproteomic studies, 79% on the identified phosphorylated residues had been localized to serines, 16% to threonines, and 5% to tyro sines, with an typical of 1. four phosphorylated residues per peptide. Gene ontology classification of the biological 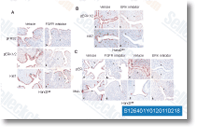 function from the 155 identified phosphoproteins indicated nearly a third of your phosphoproteins were involved in binding as their primary biological function.
function from the 155 identified phosphoproteins indicated nearly a third of your phosphoproteins were involved in binding as their primary biological function.
The PKC analogue PDBu is also a potent activator with the Brn 3b
The PKC analogue PDBu is also a potent activator of your Brn 3b promoter, and its effects can also be blocked by PD98059, suggesting that this activator converges on the p42 p44 MAPK ERK1 pathway to stimulate Brn 3b promoter activity. Dominant unfavorable MEK also blocked endogenous Brn 3b promoter activity, in a man ner that is definitely similar towards the ERK1 inhibitor, PD98059. Therefore it would appear that the p42 p44 MAPK ERK pathway is pivotal for activating the Brn 3b promoter and therefore expression in breast cancer cells. As well as stimulation by growth components, the Brn 3b promoter is strongly activated by the hormone estra diol, which regulates the growth and proliferation of typical breast epithelium as well as breast cancer cells and is significant within the etiology of breast cancer.
Oestrogens can regulate gene transcription by acting by means of one of two receptors, ERa or ERb. Our final results show that overexpression of ERa but not ERb could strongly stimulate Brn 3b promoter activity. ERa is par ticularly relevant for the improvement and progression of breast cancers since it is overexpressed in selleck a signifi cant proportion of breast cancers. In addition, ER good breast cancers are usually treated making use of recep tor antagonists, as an example, tamoxifen, as a very first line of therapy aimed at blocking ER mediated proliferative effects. Hence, the ability of ERa to stimulate Brn 3b suggests that the proliferative effects of high ER levels can be associated with the capability of ERa to trans activate other regulators, including Brn 3b, which in turn can modulate genes related with growth in these cancer cells either alone or by cooperating with ERa.
The complexity underlying the regulation with the Brn 3b promoter is increased by autoregulation, whereby Brn 3b can weakly stimulate its personal expression by bind ing to recognition sequences present in its promoter. Even so, cooperation in between Brn 3b and ERa could further boost promoter activity. Such cooperation in between selleck chemicals P276-00 Brn 3b and ERa to improve gene expression was previously observed on other ERE containing target promoters, by way of example, HSP27, exactly where Brn 3b stimu lates expression directly by binding to certain web pages inside the promoter or indirectly by interacting and cooperat ing with ER to maximally activate this promoter.
This capacity of Brn 3b to cooperate with ERa to boost gene expression, like its own, is clearly relevant to breast cancer mainly because ER expressing tumours which can be responsive to estradiol will stimulate Brn 3b, which can cooperate with ERa to additional improve its personal expression. Interestingly, mutation of  the putative ERE did not avoid ER mediated promoter activation when coexpressed with Brn 3b, but mutation in the nearby Brn three web site abolished activation by ER and its cooperation with Brn 3b.
the putative ERE did not avoid ER mediated promoter activation when coexpressed with Brn 3b, but mutation in the nearby Brn three web site abolished activation by ER and its cooperation with Brn 3b.
Tumor prone mice had been visually inspected each and every 2 day
Tumor prone mice had been visually inspected each and every two days for tumors due to the fulminant onset in these models. Tumor staging was based upon a previously described adaptation with the Intergroup Rhabdomyosarcoma Study Group staging technique. Human subjects The Oregon Well being Science University institutional re view board has produced a determination that the use of de identified tumor samples from the Nationwide Childrens Hospital Biopathology Center or Childrens Oncology Group Biorepository just isn’t human subject research due to the fact these activities do not meet the definition of human topic per 45 CFR 46. 102. Survival analysis Kaplan Meier survival analysis with the mice was performed using the endpoint becoming the development of RMS. The log rank test was utilized to ascertain the statistical sig nificance.
Each analyses were performed with Systat12 computer software. RNA isolation and quantitative reverse transcription polymerase chain reaction RNA was isolated from mouse tumors and wildtype vastus lateralis skeletal muscle using Trizol following the suppliers guidelines. RNA was then processed by RNAeasy Mini Kit and was reverse transcribed using a initial strand cDNA synthesis kit. selleck inhibitor For Figure 1A, qRT PCR analyses have been performed on an ABI7700 instrument by a Taqman assay for mouse Pax3,Foxo1a expression. The mean of 3 experimental replicates per specimen was used to calculate the ratio of gene of inter est Gapdh expression for the Taqman assay, as described previously. For Figure 1B, qRT PCR was performed making use of a regular 96 properly assay or custom Format 24 Taq man arrays applying mouse or human GAPDH as a manage for relative gene expression, and 18S RNA as a high-quality control.
Statistical considerations for this format assay have been previously described. Probesets for mouse samples were 18S Hs99999901 s1, GAPDH Mm99999915 g1, myog Mm00 446194 m1, Cdh3 Mm01249209 m1, MYCN Mm006271 79 m1, EGFR Mm00433023 m1, Fbn2 Mm00515742 m1, tcfap2b Mm00493468 m1, Hmga2 g1 and Rb1 Mm00485586 m1. Histology and immunohistochemistry Tissues buy Palbociclib fixed in 10% buffered formalin had been paraffin embedded and sectioned at three. five um thickness. Paraffin sections have been stained with hematoxylin and eosin or by Gomori Trichrome. For MyoD and Myogenin immuno histochemistry, staining was performed applying the M. O. M. Immunodection Kit Staining Procedure following the companies directions utilizing antigen unmasking.
The myogenin monoclonal major antibody was employed at a concentration of 1,50.  The Desmin monoclonal principal antibody was applied at a concentration of 1,200. For histology, we evalu ated 24 Pax3,Foxo1a,p53,Rb1 tumors, six Myf6Cre,Pax3, Foxo1a,Rb1 tumors and two Myf6Cre,Rb1 tumors. For the tissue microarray obtained in the Childrens Oncology Group Bioreposi tory, the section was pretreated with Cell Conditioning 1 for 64 minutes as antigen retrieval after which stained with rabbit polyclonal anti phospho pRb at a dilution of 1,200 followed by staining on a Ventana ES auto stainer and 3,3 diaminobenzidine detection.
The Desmin monoclonal principal antibody was applied at a concentration of 1,200. For histology, we evalu ated 24 Pax3,Foxo1a,p53,Rb1 tumors, six Myf6Cre,Pax3, Foxo1a,Rb1 tumors and two Myf6Cre,Rb1 tumors. For the tissue microarray obtained in the Childrens Oncology Group Bioreposi tory, the section was pretreated with Cell Conditioning 1 for 64 minutes as antigen retrieval after which stained with rabbit polyclonal anti phospho pRb at a dilution of 1,200 followed by staining on a Ventana ES auto stainer and 3,3 diaminobenzidine detection.
Conclusions Sialotranscriptomes of hematophagous insects have rev
Conclusions Sialotranscriptomes of hematophagous insects have revealed a large number of putative novel proteins, assistance ing to know the function of saliva in blood feeding, sugar feeding, and transmission of distinct parasites. Within the final 2 years, two black fly sialotrancriptomes have been described. The sialome of S. guianense represented the very first from a species with confirmed vectorial status for onchocerciasis. Black flies had their origin 180 MYA, primarily based on the fossil record, and presently are among the best studied Diptera, with two,025 species named, 12 of which are fossil. Their blood feeding mode has been proposed as a plesiomorphic character in the Culicomorpha appearing through the Triassic 250 MYA and diverging within the Late Jurassic.
Primarily based on tectonic plate movement, we think that Neo tropical black flies share a distant widespread origin with Neartic species, for the reason that union from the Americas only occurred during the Cenozoic, soon after the irradiation of inhibitor OSU-03012 mammals. Hence, it is actually probable that this typical black fly ancestor originated ahead of the irradiation and expan sion of mammals 60 MYA and almost certainly had birds or reptiles as their blood supply, and this origin has indeed been maintained in some species. even so, other folks could have diverged to feeding on mammals, including humans, conferring a amount of plasticity inside the Simulidae family members. For instance, S. nigrimanum was discovered to have both feeding behaviors in different places. Conversely, S. guia nense has a higher degree of anthropophily and was incri minated as the most important vector of river blindness in the focus that contains Brazil and Venezuela.
This plasticity seen in selleck inhibitor the selection of host might be accompanied by gene duplications and speedy evolution in many protein households. Here, we performed a phylogenetic evaluation of protein families discovered within the sialomes of 3 black flies from distinct subgenera S. vittatum, S. nigri manum and S. guianense. Notice that the last two are much more closely overlapping in their qualities. It really is also vital here to clear the taxonomic status of these species, mainly since  S. nigrimanum shares the same geographic distribution as S. guianense, except for S. nigrimanum absence within the Amazon area. Cur rently, some authors group both species into the Trichodagmia subgenus of Simulium, whilebased on phylogenetic analysisothers have determined that S. guianense belong to a various subgenus, Thyrso pelma, and elevated the subgenus to genus. Independent of this taxonomic confusion, it truly is clear in the phylogenetic analysis containing the black fly species that, within the majority of situations, proteins from S. nigrimanum grouped with strong bootstrap help with those of S.
S. nigrimanum shares the same geographic distribution as S. guianense, except for S. nigrimanum absence within the Amazon area. Cur rently, some authors group both species into the Trichodagmia subgenus of Simulium, whilebased on phylogenetic analysisothers have determined that S. guianense belong to a various subgenus, Thyrso pelma, and elevated the subgenus to genus. Independent of this taxonomic confusion, it truly is clear in the phylogenetic analysis containing the black fly species that, within the majority of situations, proteins from S. nigrimanum grouped with strong bootstrap help with those of S.
Immediately after 7 days in cul ture, pericytes at 80 90% conflue
Following 7 days in cul ture, pericytes at 80 90% confluency have been applied for experi ments. RBEC cultures were maintained in RBEC medium ? containing puromycin at 37 C in a humi dified atmosphere of 5% CO2 95% air, for two days. To remove the puromycin, cells were washed 3 instances with fresh RBEC medium ? and incubated with this particular medium about the third day. On the fifth day, RBECs ordinarily reached 80 90% confluency. Primary astrocyte cultures were prepared from the cere bral cortex of a single to three day outdated Wistar rats according towards the method of McCarthy and de Vellis using a slight modification. Briefly, following removing the meninges and blood vessels, the forebrains had been minced and gently dissociated by repeated pipetting in DMEM containing 10% FBS, one hundred units mL penicillin and 100 ug mL streptomycin, and filtered via a 70 um cell strainer.
Cells have been collected by centrifugation, selleck chemicals resuspended in 10% FBS DMEM and cultured in 75 cm2 flasks within a humidified atmo sphere of 5% CO2 95% air at 37 C. Cells were fed every single two 3 days by transforming medium. Following ten 14 days in culture, floating cells and weakly connected cells on the mixed main cultured cell layer were removed by vigorous shaking on the flask. Then, astrocytes in the bottom of your culture flask were trypsinized and seeded into new culture flasks. The main cultured astrocytes have been maintained in 10% FBS DMEM. They have been grown in the humidified atmo sphere of 5% CO2 95% air at 37 C. Cells at the second or third passage have been made use of for experiments. Western blot examination Brain pericytes, astrocytes and RBECs were incubated with or without unique concentrations of TNF a at 37 C for that indicated time.
When protein kinase inhibi tors were applied, they were extra 15 min just before the application of TNF a. To evaluate the expression of TNF a receptor 1 and TNF a receptor 2 amongst brain pericytes, astrocytes and RBECs, these cells had been utilized without TNF a therapy. The culture supernatants have been collected and concentrated great post to read 60 fold using Amicon Ultra centrifugal filter devices. Cells had been scraped and lysed in phosphopro tein lysis buffer containing 1% phosphatase inhibitor cocktail 1, 1% phosphatase inhibitor cocktail two and 1% protease inhibitor cocktail. The total protein concentration in cell lysates was established using a BCA Protein assay kit.
Equivalent quantities of professional tein from every sample had been electrophoretically separated on five 20% SDS polyacrylamide gels, and after that transferred to polyvinylidene difluoride membranes. Membranes had been blocked with Blocking A single or Blocking One particular P for phosphorylated proteins. Phosphorylation of p42 p44 mitogen activated protein kinase, p38 MAPK, c Jun N terminal kinase and Akt were detected with major antibodies against phospho p42 p44 MAPK, phospho p38 MAPK, phospho JNK and phospho Akt.
